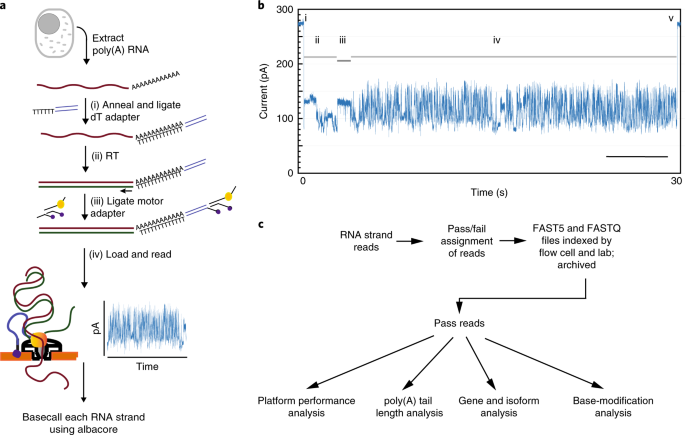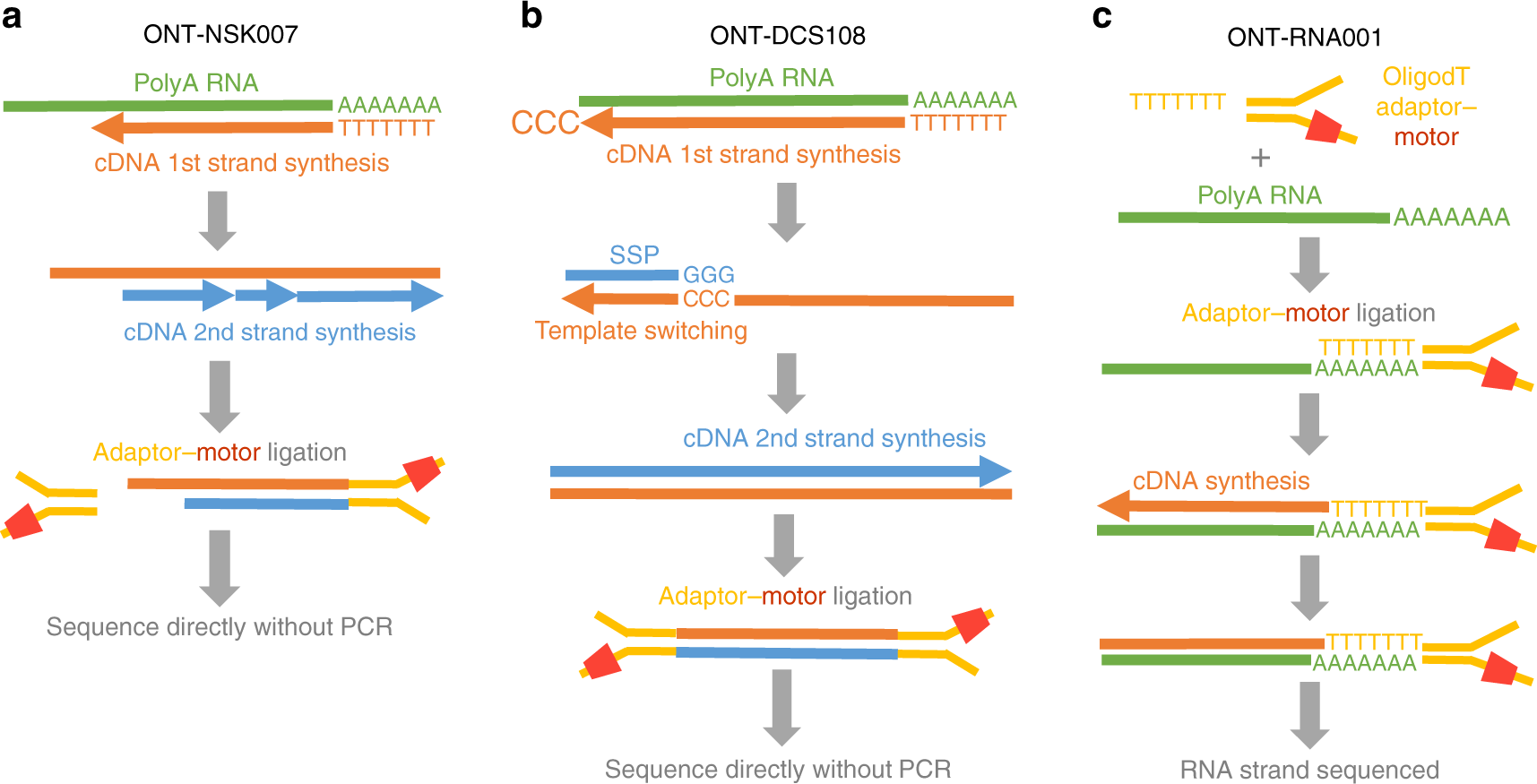
A comprehensive examination of Nanopore native RNA sequencing for characterization of complex transcriptomes | Nature Communications
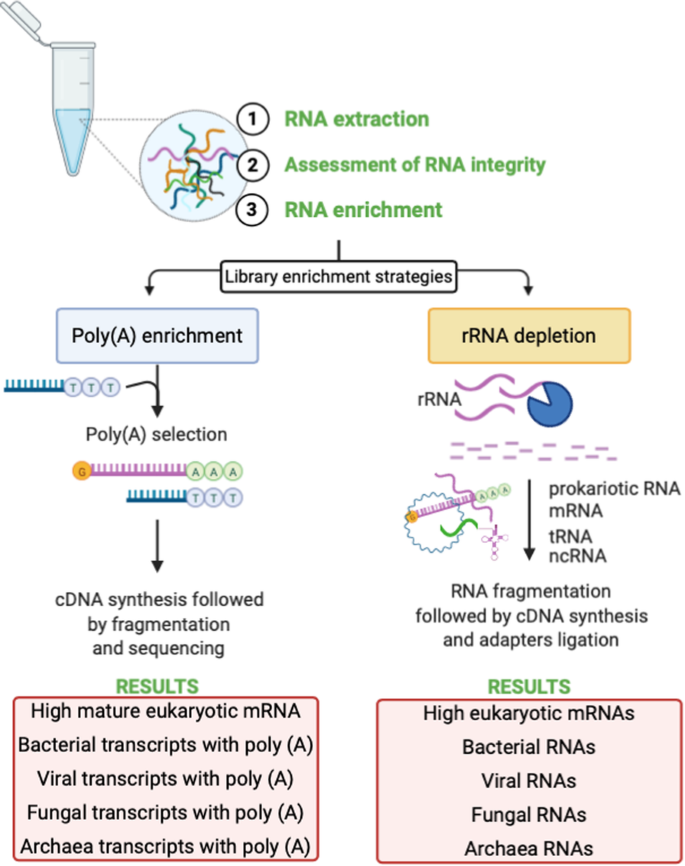
Omission of non-poly(A) viral transcripts from the tissue level atlas of the healthy human virome | BMC Biology | Full Text
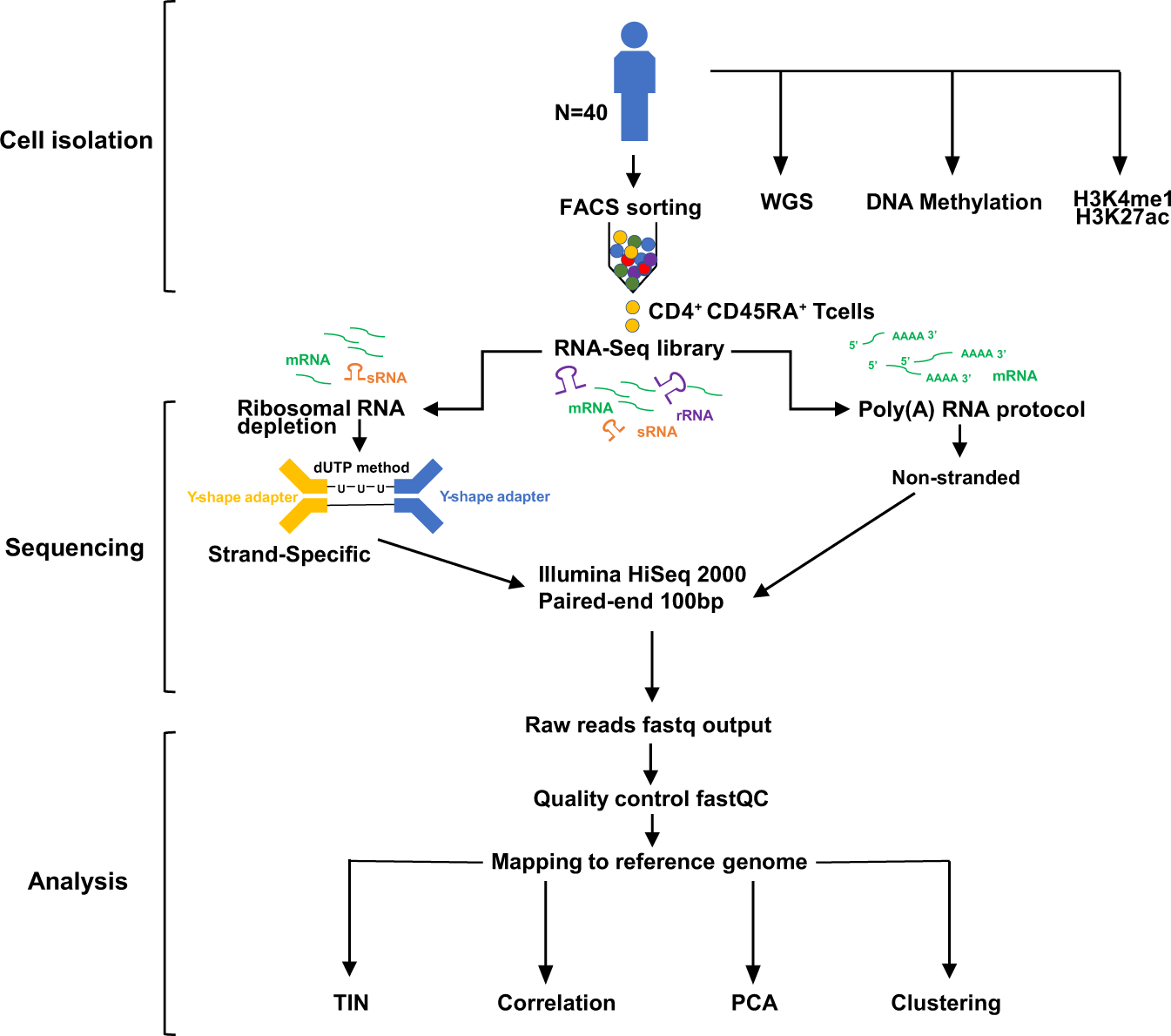
Paired rRNA-depleted and polyA-selected RNA sequencing data and supporting multi-omics data from human T cells | Scientific Data

Roles of mRNA poly(A) tails in regulation of eukaryotic gene expression | Nature Reviews Molecular Cell Biology

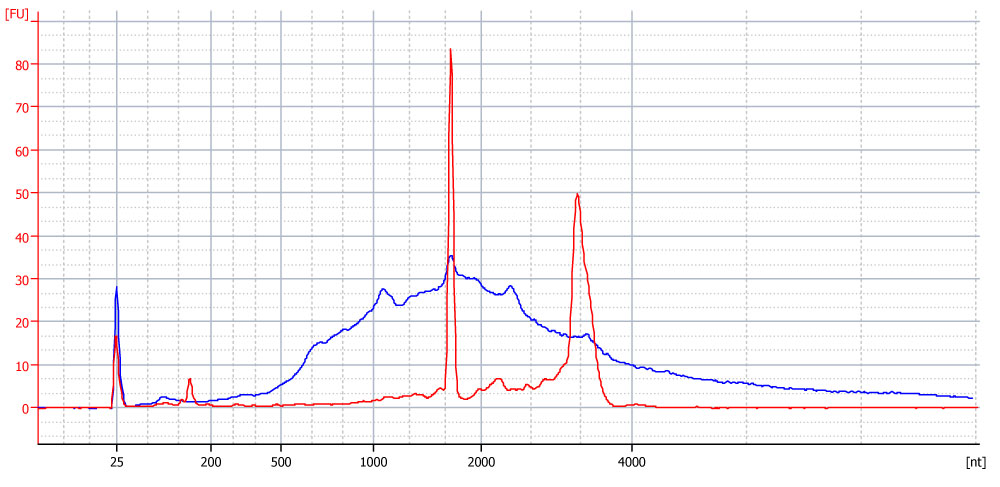
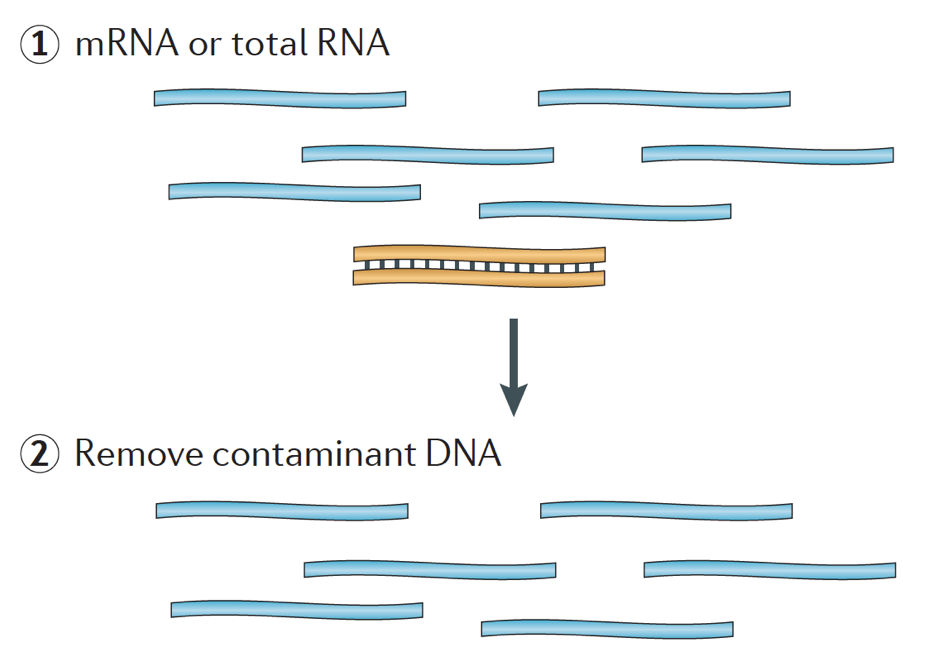
Detection of RNA extracted with different methods, Poly(A)+ RNA and the... | Download Scientific Diagram
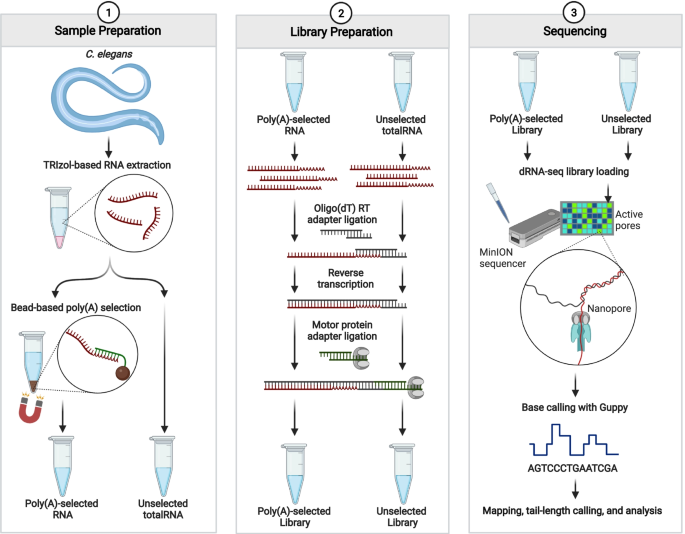
Poly(a) selection introduces bias and undue noise in direct RNA-sequencing | BMC Genomics | Full Text
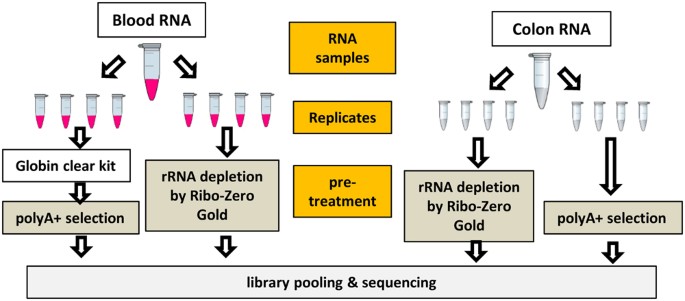
Evaluation of two main RNA-seq approaches for gene quantification in clinical RNA sequencing: polyA+ selection versus rRNA depletion | Scientific Reports

Evidence that CIP29 functions in mRNA export. (A) IF and FISH for polyA... | Download Scientific Diagram
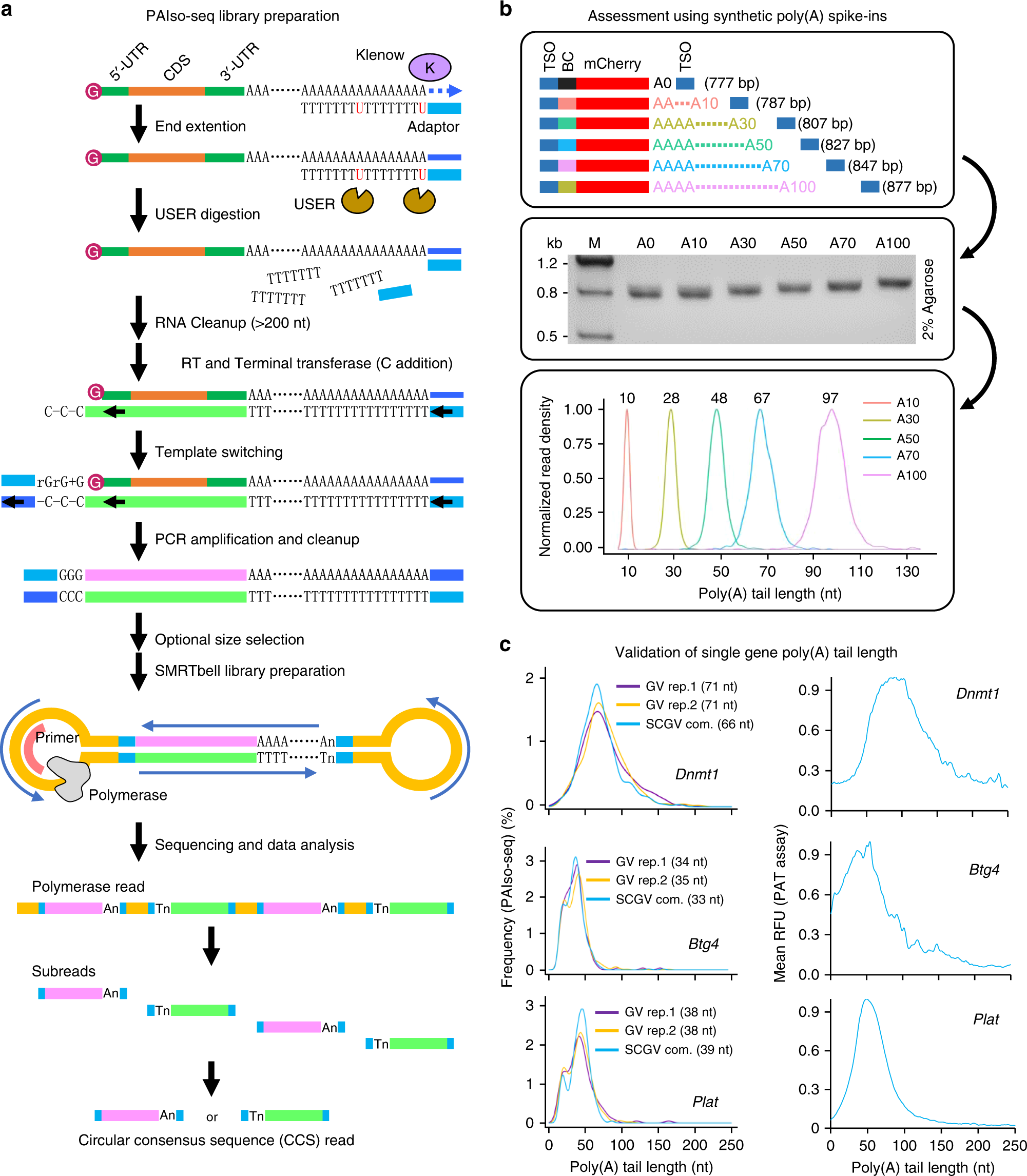
Poly(A) inclusive RNA isoform sequencing (PAIso−seq) reveals wide-spread non-adenosine residues within RNA poly(A) tails | Nature Communications
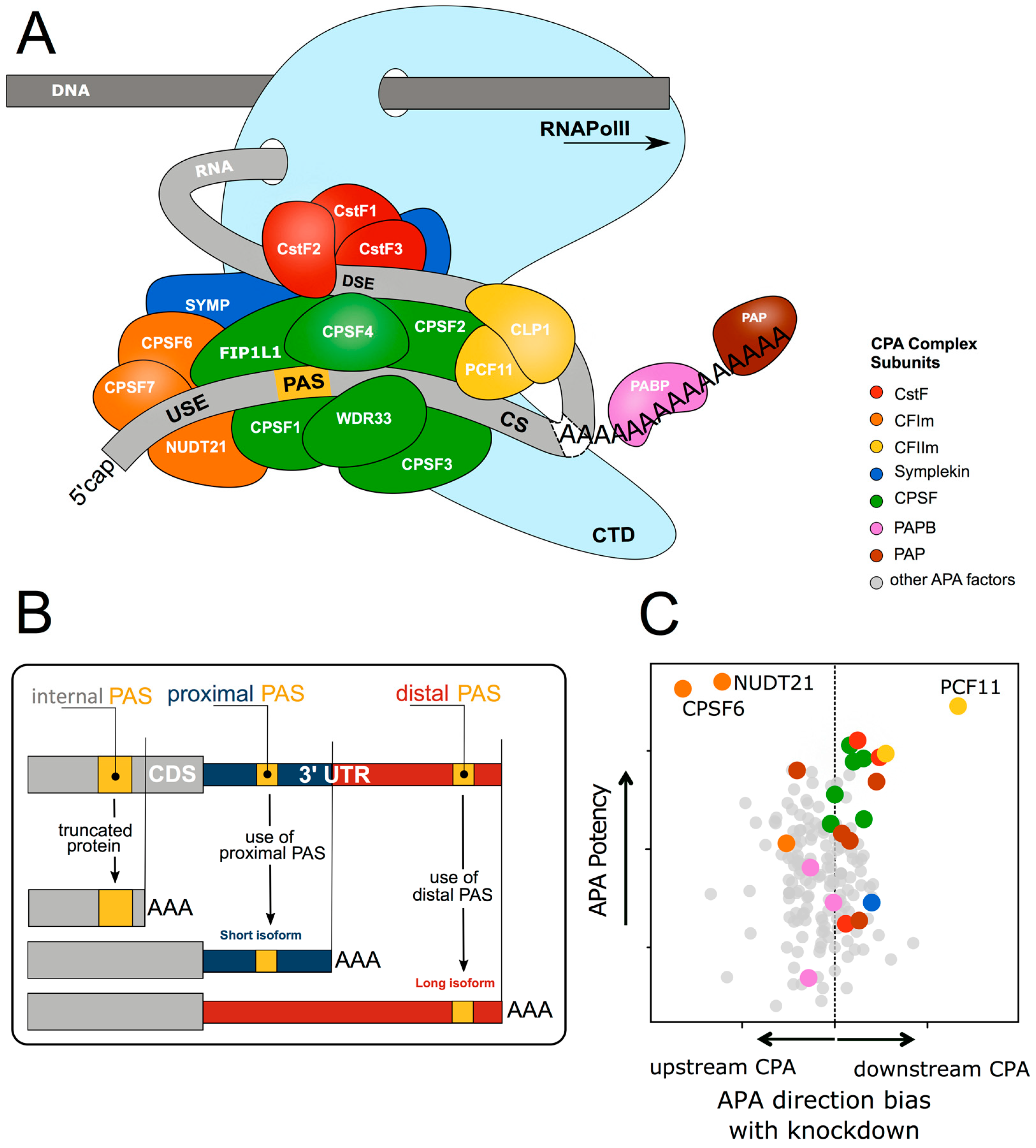
Biomolecules | Free Full-Text | Emerging Roles of RNA 3′-end Cleavage and Polyadenylation in Pathogenesis, Diagnosis and Therapy of Human Disorders